The Diversity of the DNA-Binding Landscape in the DREB/ERF Family: Focusing on Reproductive Processes in Fruit Trees With Highly Heterozygous Genome
Link to paper: https://doi.org/10.1111/pbi.70375
Published: 18 September 2025, Plant BioTechnology Journal
Summary
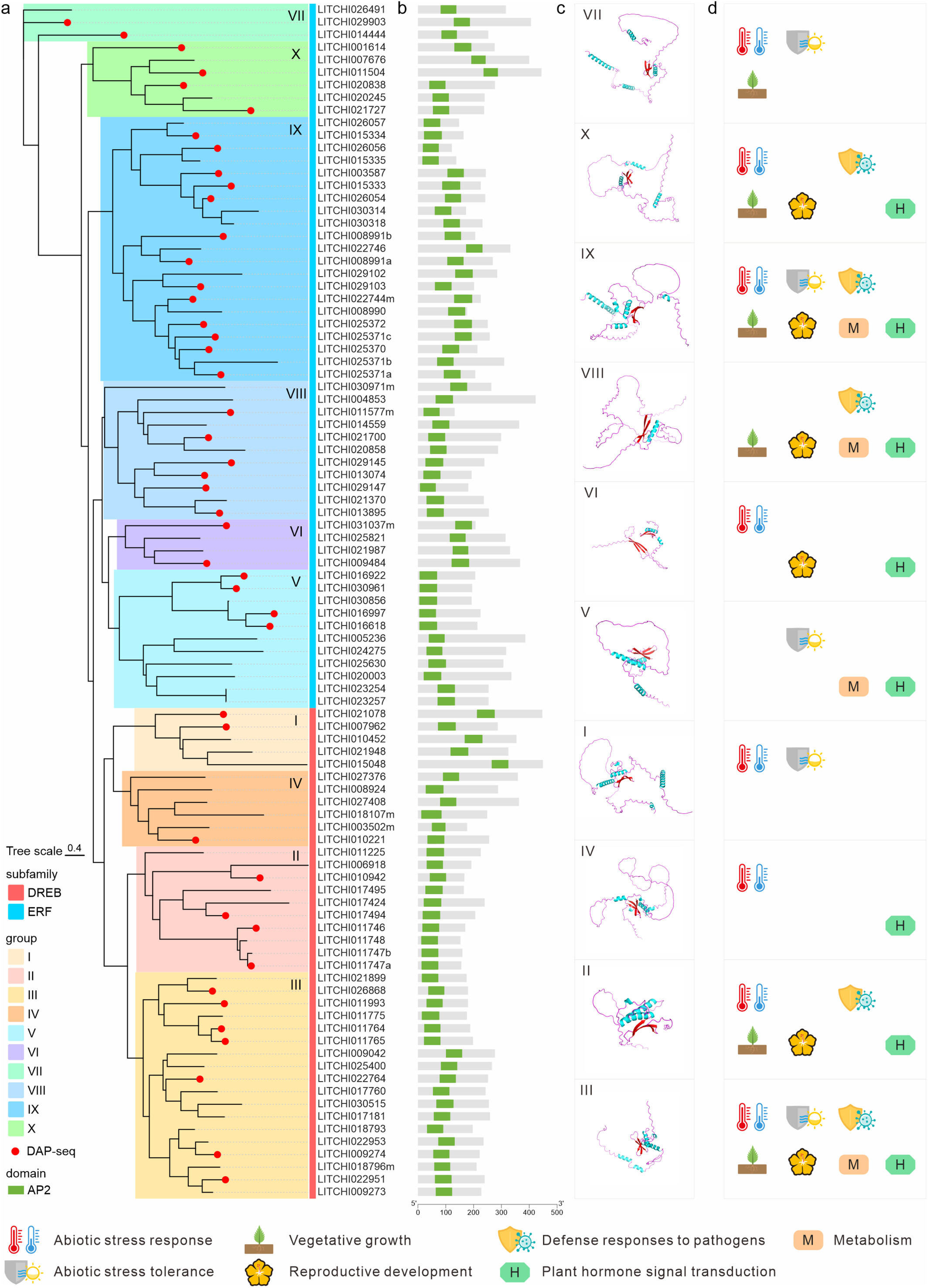
DREB/ERF transcription factors play pivotal roles in plant development; however, their structural characteristics, DNA-binding preferences, and functional roles in highly heterozygous woody plants remain insufficiently understood. Using lychee (Litchi chinensis) as a model, we identified 95 DREB/ERF genes subdivided into ten phylogenetic groups. DNA affinity purification sequencing (DAP-seq) of 45 representative members uncovered 65 194 binding sites with subfamily-specific motifs: C(G/A)CCG(A/C)C for DREB and CGCCG(C/T)C for ERF subfamilies. Each group exhibited unique binding motif preferences, aligning with their protein structures and essential peptide positions. Notably, LITCHI017494 directly regulated terpenoid biosynthesis and aroma formation by activating tandemly repeated LcTPS genes. Furthermore, single nucleotide polymorphisms (SNPs) in LITCHI017494's binding sites altered the binding efficiency of two flowering-related genes (LcSVP and LcVOZ) in early- and late-maturing haplotypes, revealing a mechanism underlying flowering and fruit maturation period. Overall, with experimental evidence, this study provides a comprehensive binding profile of the DREB/ERF family in lychee, revealing intricate transcriptional regulatory networks and serving as a crucial resource for transcription factor research within complex genomic contexts, especially in the DREB/ERF gene family.